148 total results for “”
Education
 CBCB currently supports more than 40 graduate students, mostly at the Ph.D. level. Our diverse academic community allows our students to collaborate across disciplines—machine learning researchers working with data scientists and epidemiologists, or computational biologists working with entomologists, are but two examples. Graduate students come from one of three University of Maryland graduate programs. Click the buttons below to learn more about each program.
CBCB currently supports more than 40 graduate students, mostly at the Ph.D. level. Our diverse academic community allows our students to collaborate across disciplines—machine learning researchers working with data scientists and epidemiologists, or computational biologists working with entomologists, are but two examples. Graduate students come from one of three University of Maryland graduate programs. Click the buttons below to learn more about each program.
Fall 2025 Courses
Spring 2025 Courses
CBCB Research Experiences for Undergraduates
Faculty in the CBCB regularly mentor undergraduate students in research projects. Since the creation of the Center in 2003, hundreds of undergraduate students have conducted research in the center. Many of them have pursued graduate studies and several are now faculty themselves.
Student Reseachers
CBCB currently supports more than 40 graduate students, mostly at the Ph.D. level. We also have high school interns and more than a dozen undergraduate students contributing to CBCB research. Our diverse academic community allows our students to collaborate across disciplines—machine learning researchers working with data scientists and epidemiologists, or computational biologists working with entomologists, are but two examples.
The Cummings Lab is a collaborative and dynamic lab focused on bioinformatics and computational biology, using data science, statistics, and machine learning to drive impactful research.
- Sondos Abedelhafeez
- Alexis Boleda
- Yi Chen
The El-Sayed Laboratory is focused on the study of the biology of parasitism and host-pathogen interactions using genomic approaches with the ultimate goal of better understanding infection and survival mechanisms.
- Khiem Lam
- Winnie Chen
- Amalie Ludwig
- Daniel Klimes
The Fritz Lab studies arthropod evolution in response to human-mediated environmental change. They use molecular, genomic, and computational tools to shed light on the genomic variants that facilitate adaptation.
- Theresa Menna
- Ben Gregory
- Abuzar Bhatty
- Logan Owens
- Daniel Davis
The Hall Lab studies the human gut microbiome with the goal of identifying the bacterial genes underlying health-relevant functions of the gut microbiome.
- Gabriela Arp
- Santiago Botasini
- Angela Jiang
- Sophia Levy
- Maggie R. Grant
- Robert Snell
- Owen Glass
- Alexandra Clarke
- Jonathan Yang
The research in Heng Huang's lab spans the areas of machine learning, large-scale optimization, natural language processing, computer vision, bioinformatics, and biomedical image analysis.
- Lichang Chen
- Ruibo Chen
- Yanshuo Chen
- Alireza Ganjdanesh
- Qi He
- Junyi Li
- Chenxi Liu
- Georgios Milis
- Reza Shirkavand
- Yan Wen
- Yihan Wu
- Tianyi Xiong
- Sheng Zhang
- Su Zhang
- Tong Zheng
- Ziyi Chen
- Zhenyi Wang
- Peiran Yu
- Junfeng Guo
- Shaocong Ma
- Yuke Li
Research in the Molloy Lab is at the intersection of computer science, statistics, and evolutionary genomics. They study algorithms and develop software.
- Junyan Dai
- Rachel Parsons
- Wenshan Wu
Rob Patro's COMBINE Lab spans many areas of computational/algorithmic genomics, but its core focus is on the development of algorithms, data structures, and statistical inference methods for analyzing high-throughput sequencing data.
- Hyeon Cho
- Yuan Gao
- Zahra Zare Jousheghani
- Noor Pratap Singh
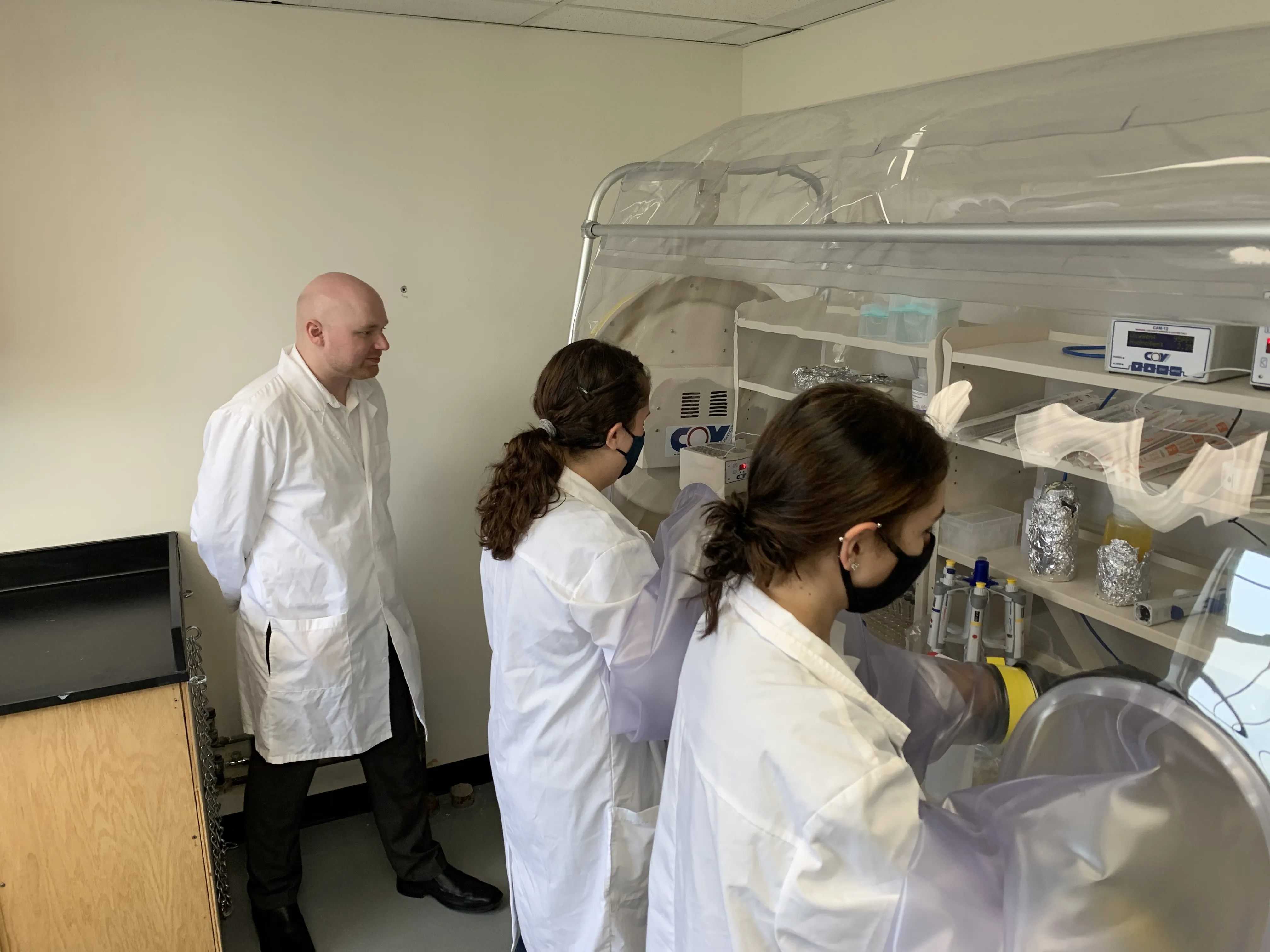





Undergraduate Research
Faculty in the CBCB regularly mentor undergraduate students in research projects. Since the creation of the Center in 2003, hundreds of undergraduate students have conducted research in the center. Many of them have pursued graduate studies and several are now faculty themselves.
We are thankful to the funders who have supported undergraduate research programs in the Center. Major funding was provided by the National Science Foundation:
Current and past undergraduate researchers in the CBCB:
2024
- Erick Alvarez (Utah Valley University), "Analysis of data on treatment for drug abuse", Mentor: Michael Cummings
- Anning Cui (University of Maryland, College Park), "Resolving containments in assembly graphs", Mentor: Mihai Pop
- Jake Jasmer (Carleton College), "Discovering new phage plasmids", Mentor: Mihai Pop
- Genelle Jenkins (Arizona State University), "Discovering new phage plasmids", Mentor: Mihai Pop
- Kashir Khan (UT Dallas), "Lineage tracing from single cell RNA-seq data", Mentor: Heng Huang
- Amara McGowan (St. Lawrence University), "Lineage tracing from single cell RNA-seq data", Mentor: Heng Huang
- James Mwihaki (MidAmerica Nazarene University), "Determining host range of bacterial plasmids", Mentor: Seth Commichaux
- Jay Rangarajan (UC San Diego), "Determining host range of bacterial plasmids", Mentor: Seth Commichaux
- Skylar Rinderknecht (Colorado School of Mines), "Indexing the CHESS transcript database", Mentor: Rob Patro
- Sophie Robertson (University of Washington), "Indexing the CHESS transcript database", Mentor: Rob Patro
- Ryan Schuenke (Drake University), "Analysis of data on treatment for drug abuse", Mentor: Michael Cummings
2023
- Vivek Agarwal (University of Maryland, College Park), "Identifying shared genomic segments across metagenomic samples", Mentor: Mihai Pop
- Sarah Blankespoor (California Polytechnic State University San Luis Obispo), "Improving sequence polishing for Nanopore reads", Mentor: Mihai Pop
- Madeline Bonanno (Tulane University), "Evaluating fast phylogenetic methods for reconstructing the evolution of SARS-CoV-2 lineages", Mentor: Erin Molloy
- John Bridgers (University of Maryland, College Park), "Improving Species Tree Estimation with Missing Data", Mentor: Erin Molloy
- Noah Cape (Williams College), “Developing a General Purpose, High Performance, Sequence pre-processor”, Mentor: Rob Patro
- Junyan Dai (University of Maryland, College Park), "Leveraging Constraints and Dynamic Programming for the Large Dollo parsimony Problem", Mentor: Erin Molloy
- Olivia Davis (Rose-Hulman Institute of Technology), “Developing an efficient algorithm for determining orthologous and paralogous relationships in a phylogeny”, Mentor: Michael Cummings
- Carola Gonzales Lebron (University of Dayton), "Evaluating multiple sequence alignment methods in light of taxonomic diversity and sampling", Mentor: Erin Molloy
- Natalia Quintana Parilla (University of Puerto Rico Mayaguez), “Investigating the splicing status of sequenced molecules from single-cell RNA-seq data”, Mentor: Rob Patro
- Charlotte Ravel (University of Maryland, College Park), "Firmware development for low-power wearable devices enabling time-tracking and low-power data transmission using Bluetooth Low Energy", Mentor: Brantley Hall
- Ashley Tien (University of Maryland, College Park), "Firmware development for low-power wearable devices enabling time-tracking and low-power data transmission using Bluetooth Low Energy", Mentor: Brantley Hall
- Gaoussou Touré (Montgomery College), “Developing an efficient algorithm for determining orthologous and paralogous relationships in a phylogeny”, Mentor: Michael Cummings
- Raunak Vijay (University of New Hampshire at Manchester), "Improving sequence polishing for Nanopore reads", Mentor: Mihai Pop
- David Zhan (University of Pennsylvania), "Firmware development for low-power wearable devices enabling time-tracking and low-power data transmission using Bluetooth Low Energy", Mentor: Brantley Hall
2022
- Esmma Almousa (University of Wisconsin Madison), "Machine learning and model interpretation for longitudinal data with applications to Parkinson's disease", Mentor: Michael Cummings
- Shria Arcot (University of California San Diego), "Improved detection of worn state in a wearable sensor", Mentor: Brantley Hall
- Nicholas Arellano (Community College of Baltimore County), "Machine learning and model interpretation for longitudinal data with applications to Parkinson's disease", Mentor: Michael Cummings
- Aditya Girish (Rutgers University), "FastMulRFS (phylogenetic inference with multi-copy genes) optimization", Mentor: Erin Molloy
- Nathan Jacobi (University of Georgia), "Single cell RNA-seq transcript quantification", Mentor: Rob Patro
- Adam Levav (University of Maryland), "Sequence alignment to detect structural variants" , Mentor: Mihai Pop
- Luiz Mata Lopez (Montgomery College), "FastMulRFS (phylogenetic inference with multi-copy genes) optimization", Mentor: Erin Molloy
- Sean Markey (University of Maryland), "Single cell RNA-seq transcript quantification", Mentor: Rob Patro
- Lucy Mettee (The University of Texas at Dallas), "Sequence alignment to detect structural variants" , Mentor: Mihai Pop
- Olivia Miskala-Dinc (University of Maryland), "Improved detection of worn state in a wearable sensor", Mentor: Brantley Hall
2021 (virtual)
- Alexander Lee (UMD), "Testing Averages for Binnacle Coverage Estimation", Mentors: Jacquelyn Michaelis, Mihai Pop
- Sho Takeshita (UMD), "Testing Averages for Binnacle Coverage Estimation", Mentors: Jacquelyn Michaelis, Mihai Pop
- Alyssa Pratt (Oregon State University), "Improving Taxonomic Classifications by Guarding Against Bad Data", Mentors: Jacquelyn Michaelis, Mihai Pop
- Celestine Zhao (UMD), "Improving Taxonomic Classifications by Guarding Against Bad Data", Mentors: Jacquelyn Michaelis, Mihai Pop
2020 (virtual)
- Shiva Mehravaran (Morgan State University), Mentors: Jacquelyn Meisel, Mihai Pop
- Chatonya Brown (Morgan State University), Mentors: Jacquelyn Meisel, Mihai Pop
- Subhadra Paudel (Morgan State University), Mentors: Jacquelyn Meisel, Mihai Pop
- Aya Kutbi (Morgan State University), Mentors: Jacquelyn Meisel, Mihai Pop
- Somayeh Moslehi (Morgan State University ), Mentors: Jacquelyn Meisel, Mihai Pop
- Christian Yovo (Towson University), Mentors: Jacquelyn Meisel, Mihai Pop
2019
- Camilo Calvo-Alcaniz (UMD), "Characterizing Synthetic Lethality Using Single- and Double-Knockout Experiments", Mentor: Sridhar Hannenhalli
- Hyeyun Chae, (Swarthmore College), "Visualizing Results from Galaxy Workflows Using Epiviz", Mentors: Jayaram Kancherla, Hector Corrada Bravo
- Marie Crane (Macalester College), "Analysis of metagenomic assembly graphs", Mentors: Jacquelyn Meisel, Mihai Pop
- Jessica Pan (Yale), "Characterizing Synthetic Lethality Using Single- and Double-Knockout Experiments", Mentor: Sridhar Hannenhalli
- Kassie Wang (high school student), "Analysis of metagenomic assembly graphs", Mentors: Jacquelyn Meisel, Mihai Pop
2018
- Larry Feldman (UMD), "CONSERVE Project and its Significant Findings", Mentor: Mihai Pop
- Kay Htut (UMD), "Application of NMF on Mutational Signatures", Mentors: Hector Corrada Bravo and Max Leiserson
- Yousef Khan (UMD), "Identification and Classification of Biothreats Case Study: Staphylococcus aureus", Mentor: Jeremy Selengut
- Nupur Pandya (UMD), "Identification and Classification of Biothreats Case Study: Staphylococcus aureus", Mentor: Jeremy Selengut
- Shilpa Roy (UMD), "Identification and Classification of Biothreats Case Study: Staphylococcus aureus", Mentor: Jeremy Selengut
- Kassie Wang (high school student), "Evaluating Refinded 16S Taxonomic Assignments", Mentor: Mihai Pop
2017
- Quinn Doubet (Wells College) - "Bacterial gene family modeling", Mentor: Jeremy Selengut
- Brian Gottfried (UMD) - "Interactive genomic visualizations", Mentor: Héctor Corrada Bravo
- Rachel Hammelman (University of Rochester)
- Ayub Khan (Virginia Commonwealth University) - "Intron retention during raspberry fruit development", Mentors: Zhongchi Liu, Sridhar Hannenhalli, Haley Wight
- Alex Lugo (UMD) - "Genetic mapping with RNA-seq data", Mentor: Steve Mount
- Reilly Thate (Rochester Institute of Technology) - "Critical assessment of metagenome interpretation", Mentors: Todd Treangen, Mihai Pop
- Michelle Tu (UMD) - "Bacterial gene family modeling", Mentor: Jeremy Selengut
2016
- Min Geol Joo (Johns Hopkins University) - "Hunt for tiny RNAs", Mentor: Sridhar Hannenhalli / Anthony Jose
- Ankita Acharya (UMD) - "Biomarkers of Healthy Human Aging", Mentor: Zia Khan
- Christopher Zawora (Georgetown University) - "Strawberry development", Mentor: Steve Mount / Zongchi Liu
- Brian Liu (UMD) - "Contamination in ancient DNA", Mentor: Philip Johnson
- Steffan Cornwell (University of Pennsylvania) - "Improving transcript quantification methods", Mentor: Steve Mount / Hector Corrada Bravo
- Marcus Fedarko (UMD) - "Metagenomic assembly", Mentor: Mihai Pop
- Morgan Walter (UMD) - "Epigenomic visualization", Mentor: Hector Corrada Bravo
2015
- Pranaya Terse (UMD) - "Comparative genomics of fruitfly to understand their development", Mentor: Zia Khan
- Kymberlee McMaster (UMD) - "Resolving haplotypes in turtle from sequencing data on multiple progeny", Mentor: Steve Mount
- Matthew Myers (UMD) - "Characterization of genomic variants in metagenomic assemblies", Mentor: Mihai Pop
- Morgan Walter (UMD) - "Protocols for interactive, collaborative, genomic visualization", Mentor: Hector Corrada-Bravo
- Teressa Chambers (University of Pittsburgh) - "Deconvolving signaling pathways from cancer data", Mentor: Sridhar Hannenhalli
2014 & Before
- Taylor Gavin (Drake University) - 2014, "Positional effects of Enhancers", Mentor: Sridhar Hannenhalli
- Michelle Hwang (University of Michigan) - 2014, "Regulatory Rewiring of engrailed gene in insects", Mentor: Leslie Pick
- Yanhong Alice Lu (UMD) - 2014, "Biosimulation of Vocal Fold Wound Healing Response to Mechanical Stress", Mentor: Nicole Li
- Kevin Nguyen (University of Maryland Baltimore County) - 2014, "Genome Assembly", Mentor: Mihai Pop
- Rama Atitkar (UMD) - 2013, "RNA-seq analysis of strawberry development", Mentor: Zhongchi Liu
- Julien Buchbinder (UMD) - 2013, "K-mer based gene segment expression analysis", Mentor: Steve Mount
- Howard Huang (Johns Hopkins) - 2013, "Assembly algorithm correctness testing", Mentors: Atif Memon, Mihai Pop
- Emily Jones (UMD) - 2013, "Evolution of Intrinsically disordered proteins", Mentor: Sridhar Hannenhalli
- Llwellyn Smith (Williams College) - 2013, "Interactive Visualization of RNA-Seq Experiments with EpiViz", Mentor: Hector Corrada-Bravo
- Mary Allison Abad (UMD) - 2012, "Motif-based and protein interaction-based sequence similarity tools", Mentor: Hannenhalli
- Elfalem Alemu (UMD) - 2012, "Epigenetic correlates of inter-tissue differences in expression variability", Mentors: Hannenhalli, Corrada-Bravo
- Jakub Bialas (UMD) - 2012, "Compression of DNA sequences from sequencing machines", Mentors: Cummings, Kingsford
- Naomi Goldstein (Wash U) - 2012, "DNA methylation genome context analysis", Mentors: Corrada-Bravo
- Michael Kleyman (UMD) - 2012, "Alternative splicing and gene expression", Mentors: Mount
- Scott Rome - 2011 - UMD - Analysis of cancer demthylation (Mentor: Hector Corrada-Bravo)
- Michele Pondlick - 2011 - UMD - (Mentor: Hector Corrada-Bravo)
- Cara Treglio - 2011 - UMD - Determination of chromatin structure from 3C data (Mentor: Carl Kingsford)
- Rachel Blythe - 2010 - Graph-based detection of genetic reassortment and rearrangement (Mentor: Carl Kingsford)
- Aashish Gadani - 2010 - Graph clustering and visualization (Mentor: Carl Kingsford)
- Nicholas Jacobs - 2010 - Gene function prediction via clustering by co-evolution (Mentor: Liliana Florea)
- JaeYoung Kwak - 2010 - Detection of variation in human genes (Mentor: Steven Salzberg)
- Joseph Paulson - 2010 - Statistics for comparing clinical data; and identification of variation in assembly graphs (Mentor: Mihai Pop)
- Megan Riordan - 2010 - Graph clustering and visualization (Mentor: Carl Kingsford)
- Joshua Wetzel - 2009 - Genome assembly complexity (Mentors: Carl Kingsford, Mihai Pop)
- Govin Vatsan (highschool)- 2009 - Visualization of assembly graphs (Mentor: Carl Kingsford)
- Petar Stojanov - 2009 - Transcriptome alignment (Mentor: Liliana Florea)
Undergraduate Research In the News
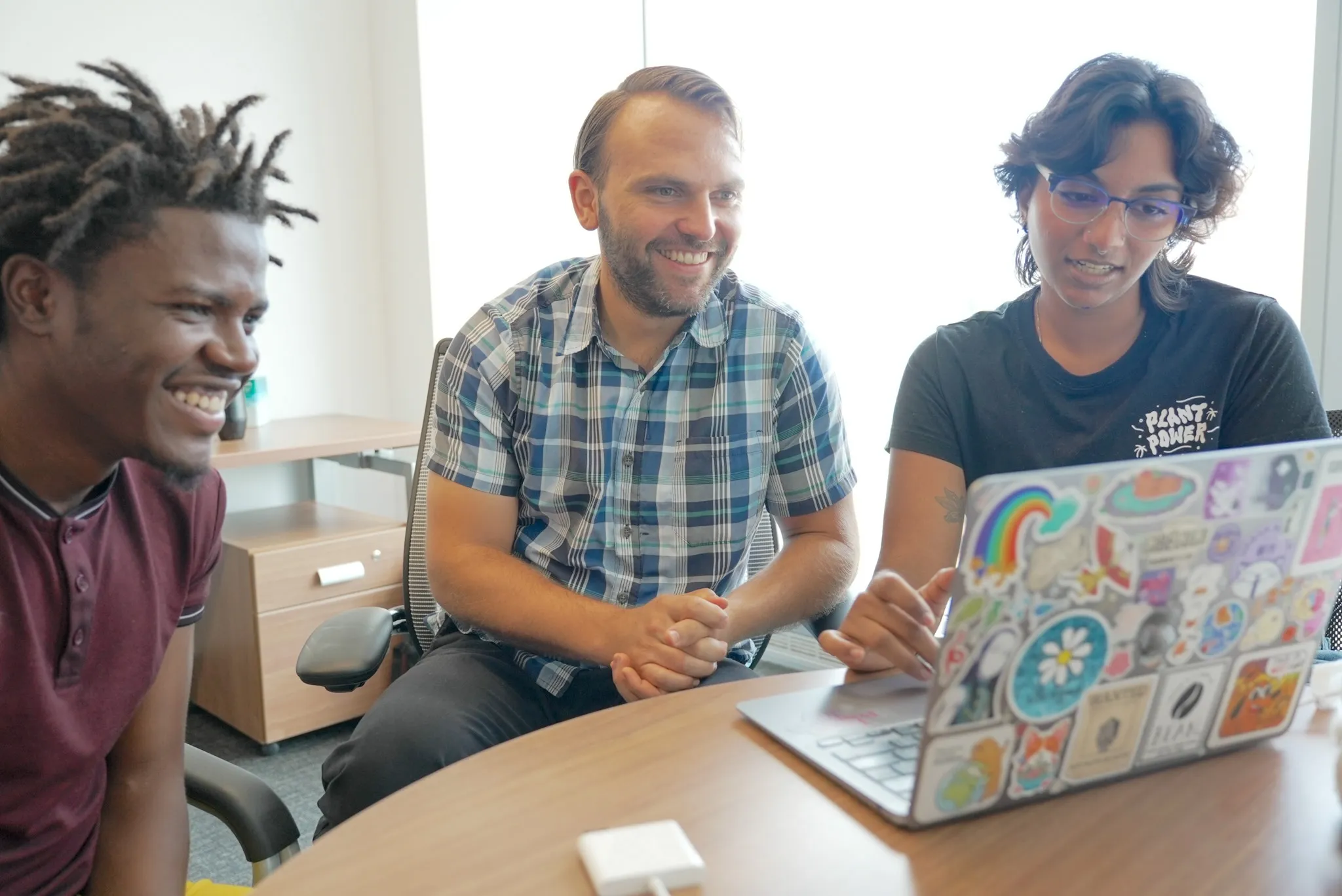

REU Bioinformatics Program Immerses Students into Interdisciplinary Research
Sponsored by NSF, the program exposes visiting undergraduates to interdisciplinary training needed to process and analyze large scale genomic datasets.

REU Interns Learn to Navigate Complexities of Health Research
From exploring the mysteries of the gut microbiome to researching the complexities of the COVID-19 virus, 14 undergraduate students from across the country…
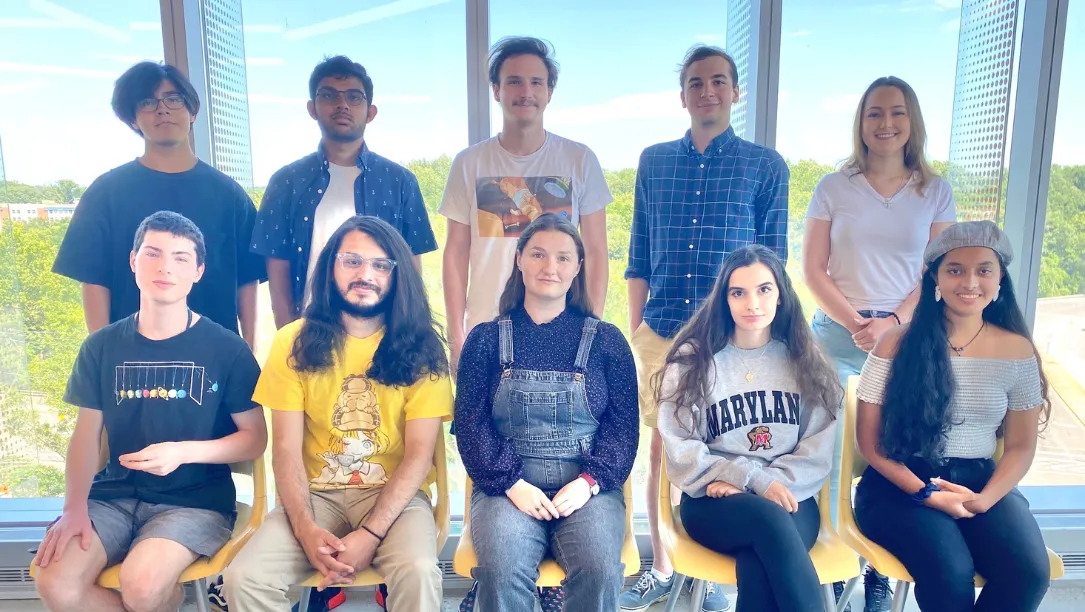

REU Summer Interns Tackle Big Problems in Bioinformatics
The 10 undergraduate students are part of an internship funded by NSF and run by faculty in CBCB.
Research
CBCB brings together researchers from many disciplines, including computer science, molecular biology, genomics, genetics, mathematics, statistics, and health computing, all of whom aspire to gain a better understanding of how life works.
We maintain robust partnerships with physicians and clinicians at the University of Maryland, Baltimore through formalized agreements like MPowering the State and the University of Maryland Institute for Health Computing.
Other research activity benefits from our close proximity to the multiple federal laboratories and agencies that dot the region, including the National Institutes of Health, the U.S. Food and Drug Administration, the National Institute of Standards and Technology, and more.

Machine Learning for Computational Medicine
Our experts use machine learning, data mining, biomedical data science, and other methods to address a broad range of biomedical problems, including Alzheimer’s disease, Parkinson’s disease, mobility issues, and more.

Parasitism and Host Pathogen Interaction
Using approaches that include the development and application of molecular, computational and phylogenetic tools, we are studying of the biology of parasitism and host-pathogen interactions using genomic approaches and other methods.

Gut Health and Nutrition
We’re exploring how research and scholarship in the microbiome sciences can result in an integrated, unifying approach to balance and optimize the health of people, animals, and the environment. This includes the use of innovative tools to measure microbial functions and improving health outcomes through microbiome modulation.
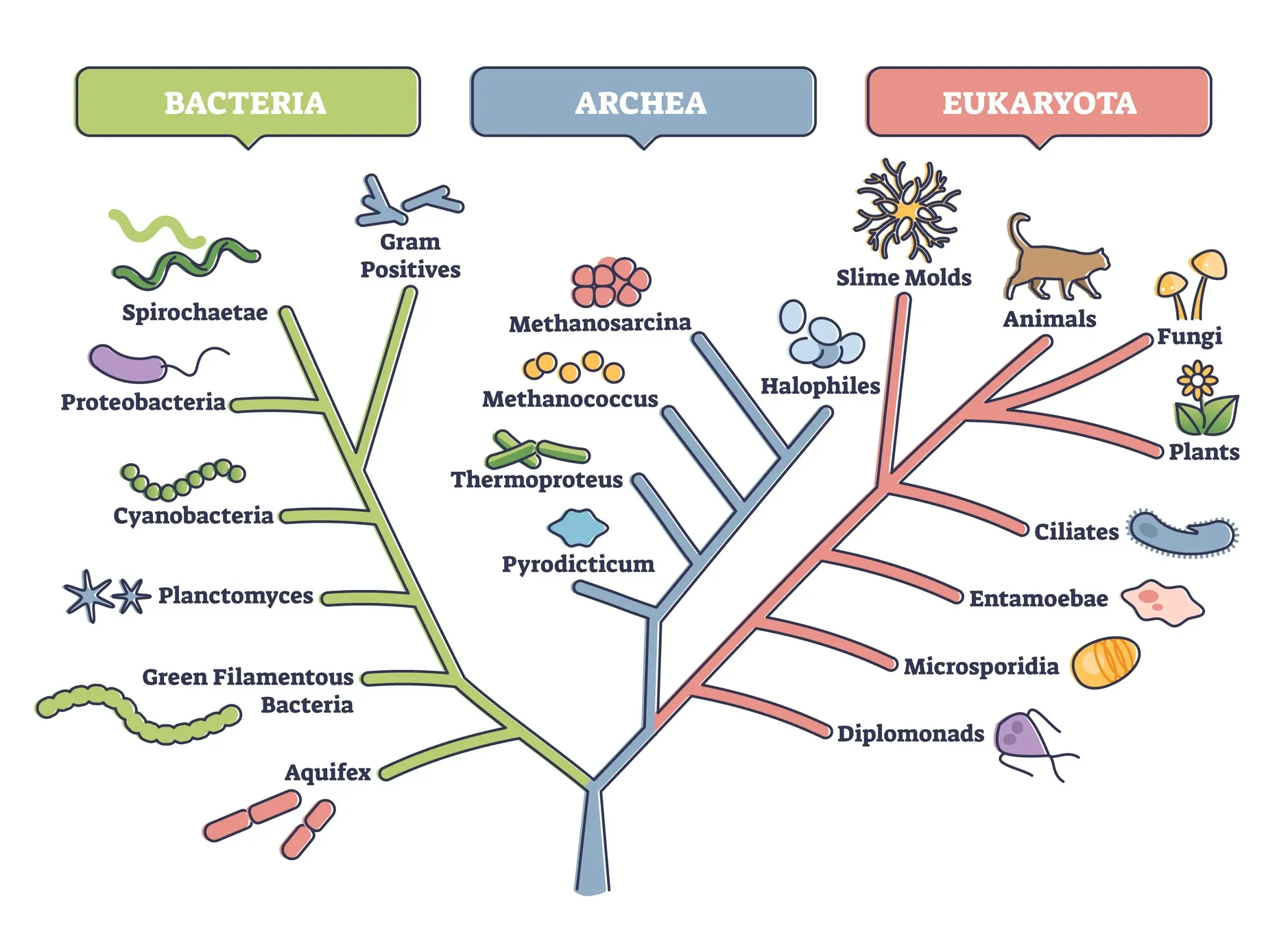
Accurate and Computationally Efficient Phylogenetic Inference
Through laboratory experiments, field sampling, and advanced molecular methods, our experts are exploring arthropod evolution in response to human-mediated environmental change and are enhancing the processing of high-throughput genome sequencing data through evolutionary modeling.
Remote Sensing to Track Disease Outbreaks
Our experts pioneered the use of bioinformatics combined with satellite imaging and other remote sensing devices to predict and monitor waterborne diseases like cholera.
Algorithms and Data Structures for Analyzing Genomic Data
We’re active in the development of computational algorithms for analyzing biological data generated through high-throughput experimental techniques, such as sequencing technologies. Our experts are also active in the design of algorithms and data structures for processing, organizing, indexing and querying high-throughput genomics data.
Accurate and Computationally Efficient Phylogenetic Inference
Through laboratory experiments, field sampling, and advanced molecular methods, our experts are exploring arthropod evolution in response to human-mediated environmental change and are enhancing the processing of high-throughput genome sequencing data through evolutionary modeling.
Our Experts:


Featured News

Developing a Genomic Method of Monitoring for Pesticide Resistance
UMIACS affiliate faculty member Megan Fritz is lead author of a study that identified genomic changes in the corn earworm—considered the costliest crop pest in…

Tracking Transgenic Crop-Produced Mutations in the Corn Earworm
Supported by a USDA grant, two UMIACS researchers have teamed up to study the corn earworm’s growing resistance to non-chemical pesticides.

Molloy Receives NSF CAREER Award to Advance Evolutionary Biology with Novel Computational Tools
The award supports the development of advanced computational methods to uncover deeper insights into species’ evolutionary relationships.
Machine Learning for Computational Medicine
Our experts use machine learning, data mining, biomedical data science, and other methods to address a broad range of biomedical problems, including Alzheimer’s disease, Parkinson’s disease, mobility issues, and more.
Our Experts:





Featured News

Cummings Awarded $2.7M to Study Impacts of Aging on the Brain
The $2.7M award will be spread across four years, supporting his team’s ongoing work that uses machine learning techniques to interpret “multiomics” data.

Cummings Employs Machine Learning to Tackle Addiction Crisis
The technology will be used to tailor treatment options to personalized patient factors like race, gender, location and educational backgrounds.
CBCB Team Publishes Innovative Research on Improving Parkinson’s Disease Diagnosis
The cross-institutional project, which uses machine learning to analyze wearable sensor data, could lead to more accurate and earlier diagnoses of people…
Parasitism and Host Pathogen Interaction
Using approaches that include the development and application of molecular, computational and phylogenetic tools, we are studying of the biology of parasitism and host-pathogen interactions using genomic approaches and other methods.
Our Expert:

Featured News

El-Sayed to Lead New Genomic Sequencing Facility at UMD
The Advanced Genomic Technologies Core, part of the Brain and Behavior Institute, will advance high-throughput and advanced large-scale genomics research, and…

$3M NIH Grant Supports Genomic Approach to Curing ‘Neglected’ Disease
Najib El-Sayed is leading a team that is using powerful sequencing tools to combat a painful, disfiguring parasitic skin condition.
UMD Researchers Discover Way to Predict Treatment Success for Parasitic Skin Disease
Findings from a new study co-authored by Najib El-Sayed could help doctors select more effective treatments earlier for patients suffering from leishmaniasis.
Remote Sensing to Track Disease Outbreaks
Our experts pioneered the use of bioinformatics combined with satellite imaging and other remote sensing devices to predict and monitor waterborne diseases like cholera.
Our Experts:


Rita R. Colwell
Featured News

Researcher Awarded $1M From NASA to Develop Disease Forecasting Center and App
Rita Colwell’s distinguished career culminates in project predicting and preventing cholera.
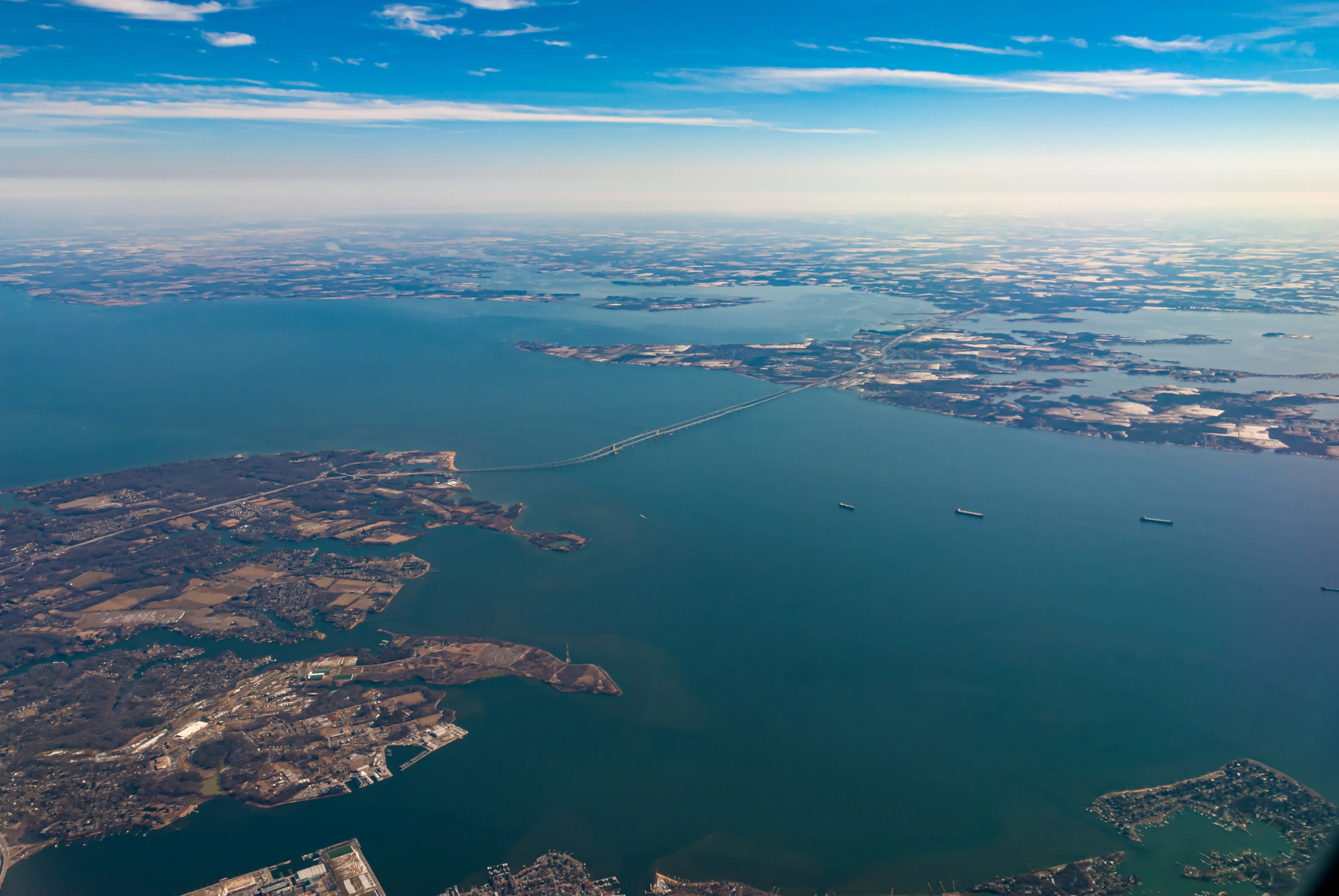

UMIACS Researchers Track the Rise of Deadly Bacteria in Maryland’s Coastal Areas
Kyle Brumfield and Rita Colwell are developing a forecasting system that can predict the risk of coming into contact with Vibrio bacteria.

Tracking Climate-Linked Pathogens Through DNA Science
UMIACS Postdoc Kyle Brumfield is using DNA sequencing and other technologies to study how environmental factors like climate can impact waterborne pathogens.
Gut Health and Nutrition
We’re exploring how research and scholarship in the microbiome sciences can result in an integrated, unifying approach to balance and optimize the health of people, animals, and the environment. This includes the use of innovative tools to measure microbial functions and improving health outcomes through microbiome modulation.
Our Experts:


Featured News

Pop Receives $2.6M NIH Grant to Develop Computational Tools for Metagenomic Analyses
The funding supports Pop's efforts to build software and develop algorithms that will help scientists better understand bacteria.

Hall Develops Device to Measure Gut Microbial Metabolism
Hall has been awarded funding from the National Institutes of Health (NIH) to develop a wearable device that will potentially lead to therapeutic interventions…

Hall Receives $3.5M in Federal Funding for Innovative Gut Microbiome Research
The funding supports groundbreaking work in the characterization of microbial enzymes and the development of novel wearable devices for monitoring gut health.
Algorithms and Data Structures for Analyzing Genomic Data
We’re active in the development of computational algorithms for analyzing biological data generated through high-throughput experimental techniques, such as sequencing technologies. Our experts are also active in the design of algorithms and data structures for processing, organizing, indexing and querying high-throughput genomics data.
Our Experts:

Rita R. Colwell




Featured News

Pop Receives $2.6M NIH Grant to Develop Computational Tools for Metagenomic Analyses
The funding supports Pop's efforts to build software and develop algorithms that will help scientists better understand bacteria.

Patro Awarded $350K to Improve Genomics Processing Tools
He will use the funding from the Chan Zuckerberg Initiative to improve upon a “constellation” of inter-related tools his lab has developed to process genomic…

Developing Advanced Tools to Improve Accuracy and Accessibility for Long-Read RNA Sequencing
Rob Patro has secured a grant from NIH to enhance computational methods for long-read RNA sequencing, technology that is crucial for investigating disease…
